Summary
- Enrichment method: GSEA
- Organism: hsapiens
-
Enrichment Categories: geneontology_Cellular_Component_noRedundant
- Interesting list: cs16_cs18_deseq2.rnk. ID type: ensembl_gene_id
- The interesting list contains 26330 user IDs in which 18841 user IDs are unambiguously mapped to 18740 unique entrezgene IDs and 7489 user IDs can not be mapped to any entrezgene ID.
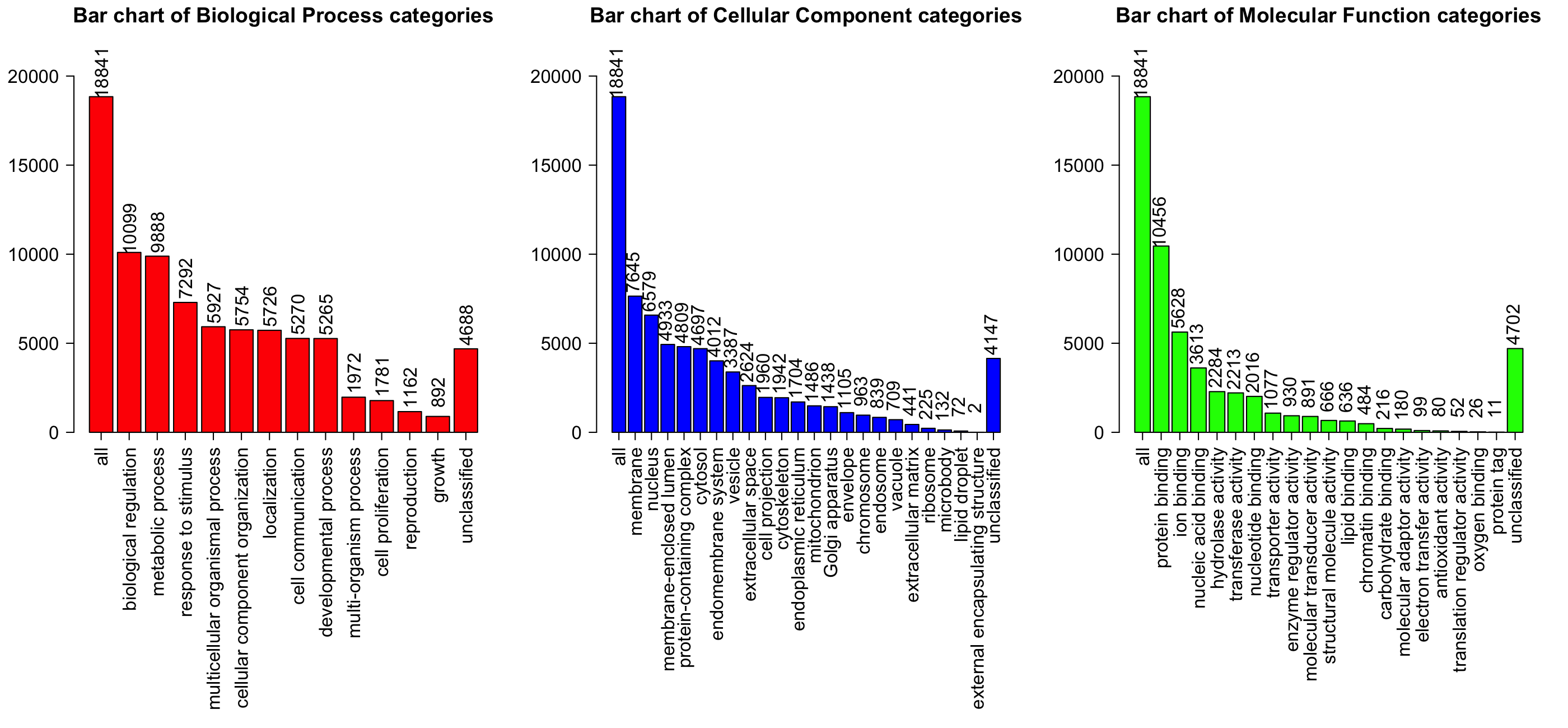
- The GO Slim summary are based upon the 18740 unique entrezgene IDs.
- Among the 18740 unique entrezgene IDs, 9157 IDs are annotated to the selected functional categories, which are used for the enrichment analysis.
Parameters for the enrichment analysis:
- Minimum number of IDs in the category: 20
- Maximum number of IDs in the category: 500
- Significance Level: FDR < 0.05
- Number of permutation: 1000
Based on the above parameters, 33 positive related categories and 19 negative related categories are identified as enriched categories , in which 50 most significant categories and represetatives in the reduced sets are shown in this report. All significant categories can be downloaded from the 'Result Download' link.
Enrichment Results